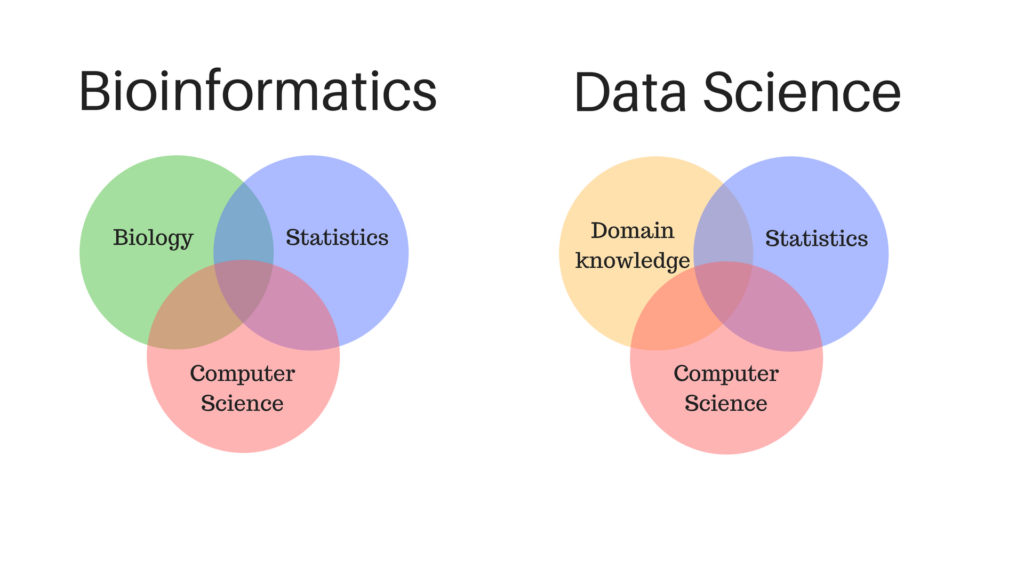
Bioinformatics has come a long way since its inception, playing a significant role in the understanding and interpretation of biological data. Often considered a relatively recent development, the beginnings of bioinformatics can be traced back to over 50 years ago when DNA analysis and the concept of desktop computers were still in their infancy. As an interdisciplinary field, bioinformatics seamlessly integrates various scientific domains like biology, chemistry, computer science, information engineering, mathematics, and statistics, enabling researchers to tackle complex large-scale data and bolster advancements in various areas of scientific research.
Initially focusing on the application of computational methods to protein sequence analysis, bioinformatics has evolved in pace with rapid technological progress and growing demands in biological research. Today, it’s primarily used to analyze biological data, develop methodologies, and design software tools for understanding myriad aspects of biology, like genomics, proteomics, and metabolomics. As a result, the world of bioinformatics has dramatically transformed how biology and information technology interact, making it increasingly difficult to distinguish the two disciplines.
The history of bioinformatics is a testament to human curiosity and the drive to better comprehend the complexity of life. With the ever-increasing availability of new biological data, the field of bioinformatics continues to push the boundaries of what’s possible, influencing multiple areas of scientific research including biomedicine, genetics, and even agriculture. As bioinformatics further develops, it will undoubtedly play an even more vital role in understanding the underpinnings of life and devising innovative solutions to pressing global challenges.
Foundations of Bioinformatics
Molecular Biology and Chemistry
The foundations of bioinformatics have strong roots in molecular biology and chemistry. In the early 1960s, researchers began applying computational methods to protein sequence analysis, including de novo sequence assembly and biological sequence databases 1. The rapid increase in data from protein biochemistry facilitated the need for advanced computational approaches to handle the growing complexity of biological systems and molecular interactions.
Mathematics and Statistics
Mathematics and statistics have always played a crucial role in the development of bioinformatics. The analysis of biological sequences, such as DNA, RNA, and protein sequences, often involves probabilistic models and statistical frameworks. Substitution models, which estimate the likelihood of one nucleotide or amino acid being replaced by another, were among the early mathematical tools applied to molecular sequences 2. Over time, advanced statistical methods have been developed to tackle diverse tasks, such as determining gene expression patterns, annotating genomic features, and inferring evolutionary relationships.
Computer Science and Engineering
Computer science and engineering have been essential in shaping the field of bioinformatics. As molecular data grew exponentially during the 1960s, it became clear that traditional manual analysis methods were insufficient 3. The advent of computers allowed researchers to analyze large datasets, develop sophisticated algorithms, and create biological databases to store and share information.
From its inception, bioinformatics has been an interdisciplinary field, relying on the synergy between molecular biology, chemistry, mathematics, statistics, computer science, and engineering. This combination of disciplines has enabled the development of powerful computational tools and methods, playing a vital role in advancing our understanding of living systems.
Early History and Pioneers
Paulien Hogeweg and Ben Hesper
Dutch theoretical biologists Paulien Hogeweg and Ben Hesper co-founded the field of bioinformatics in the early 1970s. They initially used the term to describe the study of information processes in biological systems, including computer simulation and mathematical analysis of various biological phenomena. Hogeweg and Hesper made several fundamental contributions to the field, including the development of cellular automata models to understand the complexity of living systems and investigate pattern formation in nature source.
Emile Zuckerkandl and Linus Pauling
Emile Zuckerkandl and Linus Pauling were among the first researchers to recognize the potential of using computational methods for studying molecular biology. In the early 1960s, they introduced the concept of the molecular clock, which is a method for estimating the evolutionary divergence time between different species based on the genetic differences in their macromolecules, such as DNA and proteins source. This concept laid the foundation for the development of computational methods to analyze and compare protein and nucleic acid sequences, which are now widely used in bioinformatics.
Needleman and Wunsch
Saul B. Needleman and Christian D. Wunsch developed the Needleman-Wunsch algorithm in 1970, which is considered one of the first and most important algorithms in bioinformatics. This algorithm is designed for global sequence alignment, a fundamental task in comparing and analyzing biological sequences source. Today, this algorithm serves as the basis for many other sequence alignment algorithms and continues to play a crucial role in bioinformatics research.
In conclusion, the early history of bioinformatics has deep roots in both the biological and computational sciences, with pioneers such as Paulien Hogeweg, Ben Hesper, Emile Zuckerkandl, Linus Pauling, Needleman, and Wunsch contributing to the development of its foundation. Their work has shaped the field we know today, enabling researchers to use computational tools to gain insights into the complex world of biology.
Development of Tools and Techniques
DNA and RNA Analysis
The analysis of DNA and RNA sequences has been transformed by the development of powerful computational tools. Early DNA sequencing efforts utilized the Sanger and Maxam-Gilbert methods, but these were limited by their manual nature and low throughput. With the advent of next-generation sequencing, massive amounts of DNA and RNA sequence data has become available, necessitating specialized tools and databases to manage and analyze the data.
- Sequence databases: Examples include GenBank, RefSeq, and EMBL.
- Alignment algorithms: Algorithms like Needleman-Wunsch and Smith-Waterman are used for pairwise sequence alignment, while CLUSTAL and MUSCLE are popular for multiple sequence alignment.
Popular bioinformatics software packages for DNA and RNA analysis include Biopython and Bioconductor.
Protein Sequence and Structure Analysis
Protein analysis often begins with determining the amino acid sequence of a protein, traditionally done using the Edman degradation method. Today, mass spectrometry-based methods are widely used for protein identification and characterization.
- Sequence databases: UniProt is the primary resource for protein sequences and functional information.
- Structural databases: The Protein Data Bank is the primary archive for experimentally determined protein structures.
In addition to sequence analysis, the study of protein structure and function requires computational tools for structural prediction, molecular modeling, and simulation.
Molecular Modeling and Simulation
Molecular modeling aims to generate 3D structural models of macromolecules. This includes methods like homology modeling, threading, and ab initio protein structure prediction. Software packages like MODELLER and I-TASSER are popular for these tasks.
Molecular dynamics (MD) simulations are used to study the behavior of macromolecules over time, helping to understand the relationship between structure and function. MD simulation software includes GROMACS and NAMD.
Software and Algorithms
A wide variety of software and algorithms have been developed to address specific bioinformatics problems. Common tasks include:
- Phylogenetic tree construction: Tools like MEGA and PhyML are widely used for constructing phylogenetic trees from sequence data.
- Protein docking and interaction prediction: Software like HADDOCK and ZDOCK help predict protein-protein interactions and complexes.
- Gene expression analysis: Tools such as Cufflinks and DESeq2 are used for analyzing RNA-seq data and identifying differentially expressed genes.
Overall, the development of tools and techniques in bioinformatics has provided researchers with the ability to analyze increasingly complex datasets and gain deeper insights into the workings of biological systems.
Growth of Bioinformatics Databases
Bioinformatics has grown rapidly in recent years, and this expansion has led to the development of various databases catering to different aspects of biological data. In this section, we will discuss three main categories of bioinformatics databases: Sequence Databases, Protein Structure and Interaction Databases, and Genomics and Comparative Genomics Databases.
Sequence Databases
Since the early days of bioinformatics, sequence databases have been a crucial resource for researchers. These databases store biological sequence information from DNA, RNA, and proteins. Some well-known sequence databases include:
- GenBank: A comprehensive public database containing annotated nucleotide sequences from numerous organisms.
- RefSeq: A curated reference database containing high-quality sequences of various biological molecules.
- UniProt: A comprehensive and authoritative resource for protein sequence and functional information.
These databases enable researchers to analyze and compare sequences, facilitating advancements in molecular biology and genetics.
Protein Structure and Interaction Databases
Understanding the structure and function of proteins is critical for many areas of biological research, leading to the creation of databases that catalog protein structures and interactions. Key databases in this category include:
- Protein Data Bank (PDB): A comprehensive database containing experimentally-determined three-dimensional structures of biological macromolecules, primarily proteins and nucleic acids.
- InterPro: A resource that integrates information about protein families, domains, and functional sites to provide insights into the function and evolution of proteins.
- STRING: A database that aggregates known and predicted protein-protein interactions through an integrated scoring system.
Researchers rely on these databases to inform their studies on protein function, structural genomics, molecular modeling, and drug development.
Genomics and Comparative Genomics Databases
With the advent of high-throughput sequencing technologies, genomics research has exploded, leading to the development of databases related to genomics and comparative genomics. Some notable examples include:
- Ensembl: A database providing genome annotation data for a wide range of species, including vertebrates and model organisms.
- NCBI Genome: A central portal for access to genome-scale data from the National Center for Biotechnology Information (NCBI).
- KEGG: A collection of databases that integrates genomic, chemical, and functional network information to facilitate comparative genomics and system-level analysis.
These databases play a crucial role in enabling researchers to analyze, compare, and visualize genomic data for various organisms, supporting advances in comparative genomics, evolutionary biology, and functional genomics.
Applications in Various Fields
Bioinformatics has found widespread applications in various fields. These extend from medicine and health, to agriculture and environment, and even in education and training.
Medicine and Health
The field of medicine and health has greatly benefited from the advances in bioinformatics. One of the major achievements in this area is The Human Genome Project, which has contributed to our understanding of human genetics and various diseases. With bioinformatics tools, researchers are better equipped to analyze genetic data, aiding in the discovery of new drugs and personalized medicine.
Additionally, bioinformatics has played a significant role in the field of life sciences, particularly in the study of molecular biology and biochemistry. Comparisons of genomic data between species have broadened our understanding of evolutionary relationships and functional similarities among organisms.
Agriculture and Environment
Bioinformatics has also significantly impacted the fields of agriculture and the environment. It assists in the development of improved crop varieties, pest management strategies, and environmental monitoring techniques.
- Enhancing crop breeding: Genomic information facilitates the identification of specific traits and genes responsible for desirable agricultural characteristics, such as drought resistance or increased yield.
- Pest management: Analyzing the genomic data of pests and pathogens can contribute to the development of more effective and targeted pest control strategies.
- Environmental monitoring: Bioinformatics tools can assist in monitoring ecosystems, allowing for the detection of potential issues such as pollution or population decline in specific species.
Education and Training
The discipline of bioinformatics has grown rapidly, leading to an increased demand for qualified professionals in the field. Education and training are essential for equipping students and researchers with the necessary skills to excel in this emerging area of science.
- Graduate programs: Many universities have introduced graduate programs focused on bioinformatics, providing students with opportunities to learn the theoretical and practical aspects of this interdisciplinary field.
- Online resources: There are various online platforms and tools, which help teach the principles of bioinformatics and provide access to biological databases and computational tools.
- Workshops and conferences: Regular workshops and conferences help keep researchers updated about the latest trends in bioinformatics, and facilitate networking and collaboration among professionals.
In summary, the applications of bioinformatics have been transformative in fields ranging from medicine and health to agriculture and the environment. The continued focus on education and training ensures a skilled and knowledgeable workforce, capable of tackling challenges and making new discoveries in this rapidly evolving discipline.
Bioinformatics in the Era of Big Data
Data Collection, Storage, and Manipulation
In the era of big data, bioinformatics has evolved to handle massive datasets generated through various technologies like genome sequencing, molecular dynamics simulations, and other high-throughput screening methods. The data is collected and stored in specialized databases which have been created to handle these large-scale datasets.
One significant challenge in dealing with big data in bioinformatics is the storage and manipulation of information. With the increasing volume of data generated, traditional storage methods are often insufficient. Therefore, new technologies and approaches, such as cloud-based storage and distributed computing, are employed to address these challenges.
Some key aspects of data collection, storage, and manipulation include:
- High-throughput genome sequencing generating a vast amount of genomic data
- Molecular dynamics simulations producing detailed information on interactions between molecules
- Advanced data storage solutions like cloud-based systems and distributed computing to handle the growing volume of data
Big Data Analytics and Machine Learning
The unprecedented scale of biological data generated through various techniques has necessitated the development of robust bioinformatics tools to make sense of this information. Big data analytics and machine learning play a crucial role in processing and analyzing these vast datasets efficiently.
Big data analytics involves the use of specific statistical algorithms and computational methods to identify patterns and trends within the data. Machine learning, an essential component of big data analytics, enables these algorithms to adapt and improve their performance as more data becomes available.
Some critical aspects of big data analytics and machine learning in bioinformatics include:
- Utilizing machine learning algorithms to reveal hidden patterns and relationships within the data
- The application of unsupervised learning techniques, such as clustering, to group similar biological entities
- Leveraging supervised learning methods, like classification algorithms, to predict outcomes based on input data
The availability of big data in bioinformatics has indeed provided new opportunities and challenges. By developing advanced tools and techniques, researchers can now gain deeper insights into biological systems and improve our understanding of the complex world within and around us.
Current Challenges and Future Directions
Integrating Omics Technologies
The integration of various omics technologies, such as genomics, proteomics, and transcriptomics, is a significant challenge in the field of bioinformatics. These technologies generate large volumes of data, and the analysis and interpretation of this data require the development of advanced computational tools and algorithms. The National Institutes of Health (NIH) has been funding research to facilitate the integration of omics data, enabling a more comprehensive understanding of biological systems. The advancement of functional genomics and structural genomics plays a crucial role in this integration process, allowing for improved drug designing and biomarker identification.
Expanding Scope of Computational Biology
The expanding scope of computational biology encompasses a wide range of research areas, from medical informatics to protein interactions. As bioinformatics continues to evolve, the demand for skilled computational biologists who can develop innovative statistical methods and algorithms has increased. Collaborations between different fields, such as biotechnology, computer science, and mathematics, are essential to develop new approaches for data analysis and visualization. These interdisciplinary efforts are facilitated by resources such as the National Center for Biotechnology Information (NCBI) and the World Wide Web, which provide centralized repositories for biological data and enable global scientific collaboration.
Addressing Ethical and Legal Issues
As bioinformatics continues to advance, ethical and legal issues surrounding data privacy, security, and usage become increasingly important. The large-scale generation of personal genomic data raises concerns about consent, confidentiality, and the potential for discrimination based on genetic information. Addressing these issues requires the development of comprehensive guidelines and regulations that protect individual privacy while also promoting knowledge-sharing and scientific progress. Collaboration between stakeholders, including researchers, policymakers, and regulatory agencies, is essential to create a responsible and ethical environment for the future of bioinformatics.